Medea
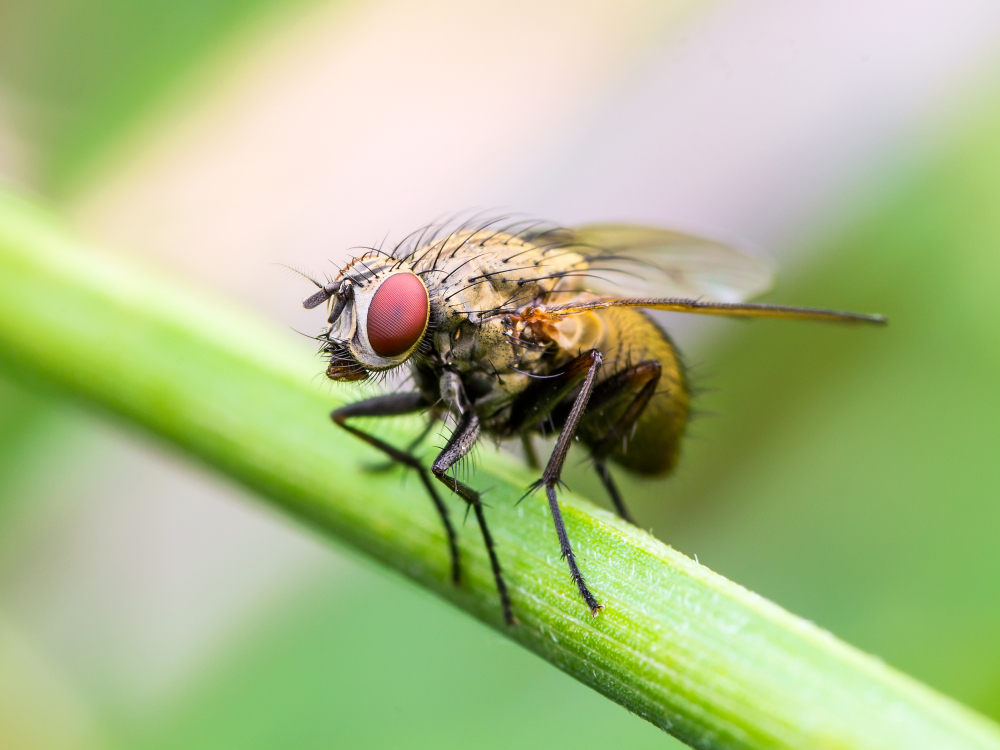
fruit fly Drosophila
Medea is a gene from the fruit fly Drosophila melanogaster that was one of the first two Smad genes discovered. For both genes, the maternal effect lethality was the basis for selection of their names. Medea was named for the mythological Greek Medea, who killed her progeny fathered by Jason.

Medea and Mothers
Both Medea and Mothers against dpp were identified in a genetic screen for maternal effect mutations that caused lethality of heterozygous decapentaplegic progeny. Because decapentaplegic is a bone morphogenetic protein in the transforming growth factor beta superfamily, identification of the fly Smad genes provided a much needed clue to understand the signal transduction pathway for this diverse family of extracellular proteins. Humans, mice, and other vertebrates have a gene with the same function as Medea, called SMAD4. An overview of the biology of Medea is found at The Interactive Fly, and the details of Medea genetics and molecular biology are curated on Flybase.
Suspendisse porttitor sollicitudin nisi et bibendum. Donec ac semper ante, non sagittis sapien. Duis hendrerit tellus sit amet cursus euismod.
Fusce a porttitor augue
Another laboratory used Medea as an acronym to describe a synthetic gene causing Maternal effect dominant embryonic arrest. The formal genetic designation for Maternal effect dominant embryonic arrest is PMedea.myd88, more details are in Flybase.
Smads are roughly between 400 and 500 amino acids long, and consist of two globular regions at the amino and carboxy termini, connected by a linker region. These globular regions are highly conserved in R-Smads and Co-Smads, and are called Mad homology 1 (MH1) at the N-terminus, and MH2 at the C-terminus. The MH2 domain is also conserved in I-Smads. The MH1 domain is primarily involved in DNA binding, while the MH2 is responsible for the interaction with other Smads and also for the recognition of transcriptional co-activators and co-repressors. R-Smads and Smad4 interact with several DNA motifs though the MH1 domain. These motifs include the CAGAC and its CAGCC variant, as well as the 5-bp consensus sequence GGC(GC)|(CG). Receptor-phosphorylated R-Smads can form homotrimers, as well as heterotrimers with Smad4 in vitro, via interactions between the MH2 domains.
Trimers of one Smad4
Trimers of one Smad4 molecule and two receptor-phosphorylated R-Smad molecules are thought to be the predominant effectors of TGF-β transcriptional regulation. The linker region between MH1 and MH2 is not just a connector, but also plays a role in protein function and regulation. Specifically, R-Smads are phosphorylated in the nucleus at the linker domain by CDK8 and 9, and these phosphorylations modulate the interaction of Smad proteins with transcriptional activators and repressors. Furthermore, after this phosphorylation step, the linker undergoes a second round of phosphorylations by GSK3, labelling Smads for their recognition by ubiquitin ligases, and targeting them for proteasome-mediated degradation. The transcription activators and the ubiquitin ligases both contain pairs of WW domains. These domains interact with the PY motif present in the R-Smad linker, as well as with the phosphorylated residues located in the proximity of the motif. Indeed, the different phosphorylation patterns generated by CDK8/9 and GSK3 define the specific interactions with either transcription activators or with ubiquitin ligases. Remarkably, the linker region has the highest concentration of amino acid differences among metazoans, although the phosphorylation sites and the PY motif are highly conserved.